This post is to illustrate how to use R to do basic conjoint analysis.
Conjoint Analysis
Marketing people used to use Sawtooth to do conjoint analysis.
What is conjoint analysis? Here is the intro from Sawtooth:
“Conjoint analysis methods use statistical analysis to compute mathematical representations of survey respondents’ preferences for product features, simulate how attribute changes affect demand, and even predict market acceptance of new products before they launch.
This powerful research methodology has become very popular with market researchers and economists for the valuable insight that few other research methods can provide.”
“Consider the task of purchasing a new vehicle.
When searching for the “right” vehicle, you typically consider a multitude of variables: type, make (brand), model, year, mileage, price, color, accessories packages, etc. You can see how this is a complex choice to make.
How does anyone possibly make such a complicated decision?
You think about what features matter most to you and then make tradeoffs.
For example, one vehicle could be an acceptable make and model, but at a high price point, whereas a second option is not the ideal make and model but has a more palatable price.
The tradeoff in this simple example is whether the make and model is more preferable to a reasonable price, or vice versa.
A conjoint survey captures the relationship between different attributes of a product and how preferable they are both comparatively and when combined.
Because of this, conjoint analysis is the gold standard solution for testing multiple features simultaneously, when limited A/B testing doesn’t go far enough.”
“In simple terms, the conjoint survey design algorithm generates a balanced survey design with product profiles, also known as concepts, formulated to have ideal statistical properties (such as level balance and independence of attributes).
Each profile is made up of multiple conjoined product features (hence, conjoint analysis), such as brand, type, engine, and color, each with systematically varied levels.
These product concepts are then included in a series of questions, usually 8-20, called choice tasks, that make up the conjoint analysis portion of the survey.”
Example
I am following the example here: https://bookdown.org/lien_lamey/R_tutorial/8-1-data-5.html
library(tidyverse)
library(openxlsx)
url <- "https://feb.kuleuven.be/lien.lamey/public/rmodule/icecream.xlsx"
icecream <- read.xlsx(url)
icecream <- icecream %>%
pivot_longer(cols = starts_with("Individual"), names_to = "respondent", values_to = "rating") %>% # respondent keeps track of the respondent, rating will store the respondent's ratings, and we want to stack every variable that starts with Individual
rename("profile" = "Observations") %>% # rename Observations to profile
mutate(profile = factor(profile), respondent = factor(respondent), # factorize identifiers
Flavor = factor(Flavor), Packaging = factor(Packaging), Light = factor(Light), Organic = factor(Organic)) # factorize the ice cream attributes
icecream## # A tibble: 150 × 7
## profile Flavor Packaging Light Organic respondent rating
## <fct> <fct> <fct> <fct> <fct> <fct> <dbl>
## 1 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 1
## 2 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 6
## 3 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 5
## 4 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 1
## 5 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 2
## 6 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 7
## 7 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 7
## 8 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 5
## 9 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 1
## 10 Profile 1 Raspberry Homemade waffle No low fat Not organic Individual… 10
## # ℹ 140 more rowsIn this example, we have 4 attributes: Flavor, Packaging, Light, and Organic. There are 5, 3, 2, 2 levels correspondingly. Usually we are interested in how much does each level of attributes predict the outcome, say the rating of the icecream. To do this, we would think we’ll need all 532*2=60 combinations of attribute levels to estimate the effect. In that case, we would need data for each combination. This is called full factorial design. The problem of full factorial design is that it might be too costly or not possible if the combination is huge. Instead, conjoint analysis allows us to estimate the effect of each level of each attribute by using a fraction of the combinations. This is done by creating a set of profiles, each of which contains a subset of the levels of the attributes. The profiles are then presented to the respondents, who rate the profiles. The ratings are then used to estimate the effect of each level of each attribute. This is done by regressing the ratings on the levels of the attributes. The coefficients of the regression are the estimates of the effects of the levels of the attributes.
The disadvantage of a fractional factorial design is that some effects are confounded; that is, the effect of one level of an attribute is confounded with the effect of another level of another attribute.
# attribute1, attribute2, etc. are vectors with one element in which we first provide the name of the attribute followed by a semi-colon and then provide all the levels of the attributes separated by semi-colons
attribute1 <- "Flavor; Raspberry; Chocolate; Strawberry; Mango; Vanilla"
attribute2 <- "Package; Homemade waffle; Cone; Pint"
attribute3 <- "Light; Low fat; No low fat"
attribute4 <- "Organic; Organic; Not organic"
# now combine these different attributes into one vector with c()
attributes <- c(attribute1, attribute2, attribute3, attribute4)
attributes## [1] "Flavor; Raspberry; Chocolate; Strawberry; Mango; Vanilla"
## [2] "Package; Homemade waffle; Cone; Pint"
## [3] "Light; Low fat; No low fat"
## [4] "Organic; Organic; Not organic"Conjoint design of experiment
The first step in conjoint analysis is to create a design of experiment. This is to use a fraction of all possible combinations of attribute levels to estimate the effect of each level of each attribute. For example, instead of using all 60 icecream profiles, we use 10 profiles.
We use “radiant” package’s “doe” function to create the design of experiment. The “doe” function takes the attributes as input and returns a data frame with the profiles. The “seed” argument is used to fix the random number generator. This is useful if you want to reproduce the same results.
library(radiant)
ice_doe <- doe(attributes, seed = 123)
ice_doe$eff## Trials D-efficiency Balanced
## 1 9 0.105 FALSE
## 2 10 0.389 FALSE
## 3 11 0.411 FALSE
## 4 12 0.614 FALSE
## 5 13 0.542 FALSE
## 6 14 0.479 FALSE
## 7 15 0.762 FALSE
## 8 16 0.738 FALSE
## 9 17 0.748 FALSE
## 10 18 0.756 FALSE
## 11 19 0.644 FALSE
## 12 20 0.895 FALSE
## 13 21 0.848 FALSE
## 14 22 0.833 FALSE
## 15 23 0.790 FALSE
## 16 24 0.827 FALSE
## 17 25 0.787 FALSE
## 18 26 0.768 FALSE
## 19 27 0.759 FALSE
## 20 28 0.736 FALSE
## 21 29 0.702 FALSE
## 22 30 0.984 TRUE
## 23 31 0.952 FALSE
## 24 32 0.933 FALSE
## 25 33 0.928 FALSE
## 26 34 0.900 FALSE
## 27 35 0.871 FALSE
## 28 36 0.893 FALSE
## 29 37 0.866 FALSE
## 30 38 0.843 FALSE
## 31 39 0.836 FALSE
## 32 40 0.922 FALSE
## 33 41 0.899 FALSE
## 34 42 0.904 FALSE
## 35 43 0.882 FALSE
## 36 44 0.861 FALSE
## 37 45 0.949 FALSE
## 38 46 0.919 FALSE
## 39 47 0.912 FALSE
## 40 48 0.911 FALSE
## 41 49 0.891 FALSE
## 42 50 0.959 FALSE
## 43 51 0.939 FALSE
## 44 52 0.944 FALSE
## 45 53 0.925 FALSE
## 46 54 0.924 FALSE
## 47 55 0.906 FALSE
## 48 56 0.902 FALSE
## 49 57 0.884 FALSE
## 50 58 0.872 FALSE
## 51 59 0.855 FALSE
## 52 60 1.000 TRUE#summary(doe(attributes, seed = 123, trials = 10))This is a list of efficiency of different number of “trials”. Trials mean combinations of attribute levels. For example, with 10 trials, the efficiency is .389 and it’s unbalanced.
We can see that the only two designs with balanced design are 30 and 60. That’s because 30 and 60 are the only two numbers that can be divided by 5, 3 and 2, all unique number of 5, 3, 3, 2. If the factors are, say, 3, 2, 3. Then the integers that can be divided by 3 and 2 are 6, 12, and 18.
Efficiency is a measure of the information content a design can capture. Efficiency are typically stated in relative terms, as in “design A is 80% as efficient as design B.” In practical terms this means you will need 25% more (the reciprocal of 80%) design A observations (respondents, choice sets per respondent or a combination of both) to get the same standard errors and significance as with the more efficient design B. We used a measure of efficiency called D-Efficiency (Kuhfeld et al. 1994). In this case, the only orthogonal design is the full factorial design of 60 trials.
For example, a trial of 30 has a efficiency of .984, that means it’s 98.4% as efficient as the full factorial design of 60 trials.
library(radiant)
ice_doe2 <- doe(attributes, seed = 123, trials = 30)
ice_doe2$cor_mat## Flavor Package Light Organic
## Flavor 1.000000e+00 -1.942890e-16 -1.665335e-16 -1.665335e-16
## Package -1.942890e-16 1.000000e+00 -1.665335e-16 -1.665335e-16
## Light -1.665335e-16 -1.665335e-16 1.000000e+00 -1.045299e-01
## Organic -1.665335e-16 -1.665335e-16 -1.045299e-01 1.000000e+00From the covariance matrix, we can se that in this 30 profile design, there is a correlation of -.105 between “Light” and “Organic”. That means these two effects are confounded; the design is not orthogonal. That’s why the efficiency is not 1. The D-efficiency is calculated as a function of the determinant of the correlation matrix. The D-efficiency of the full factorial design is 1. The D-efficiency of the 30 profile design is .984. That means the 30 profile design is 98.4% as efficient as the full factorial design.
The partial factorial design of 30 profile is
ice_doe2$part## trial Flavor Package Light Organic
## 1 1 Raspberry Homemade_waffle Low_fat Organic
## 2 4 Raspberry Homemade_waffle No_low_fat Not_organic
## 3 6 Raspberry Cone Low_fat Not_organic
## 4 7 Raspberry Cone No_low_fat Organic
## 5 10 Raspberry Pint Low_fat Not_organic
## 6 11 Raspberry Pint No_low_fat Organic
## 7 13 Chocolate Homemade_waffle Low_fat Organic
## 8 14 Chocolate Homemade_waffle Low_fat Not_organic
## 9 19 Chocolate Cone No_low_fat Organic
## 10 20 Chocolate Cone No_low_fat Not_organic
## 11 22 Chocolate Pint Low_fat Not_organic
## 12 23 Chocolate Pint No_low_fat Organic
## 13 26 Strawberry Homemade_waffle Low_fat Not_organic
## 14 28 Strawberry Homemade_waffle No_low_fat Not_organic
## 15 29 Strawberry Cone Low_fat Organic
## 16 32 Strawberry Cone No_low_fat Not_organic
## 17 33 Strawberry Pint Low_fat Organic
## 18 35 Strawberry Pint No_low_fat Organic
## 19 39 Mango Homemade_waffle No_low_fat Organic
## 20 40 Mango Homemade_waffle No_low_fat Not_organic
## 21 41 Mango Cone Low_fat Organic
## 22 42 Mango Cone Low_fat Not_organic
## 23 46 Mango Pint Low_fat Not_organic
## 24 47 Mango Pint No_low_fat Organic
## 25 49 Vanilla Homemade_waffle Low_fat Organic
## 26 51 Vanilla Homemade_waffle No_low_fat Organic
## 27 53 Vanilla Cone Low_fat Organic
## 28 56 Vanilla Cone No_low_fat Not_organic
## 29 58 Vanilla Pint Low_fat Not_organic
## 30 60 Vanilla Pint No_low_fat Not_organicConjoint analysis
In the icecream data set we read in, it’s a 10 profile design. There are 15 respondents. They rate 10 profiles of icecreams.
conjoint <- conjoint(icecream, rvar = "rating", evar = c("Flavor","Packaging","Light","Organic"))
summary(conjoint)## Conjoint analysis
## Data : icecream
## Response variable : rating
## Explanatory variables: Flavor, Packaging, Light, Organic
##
## Conjoint part-worths:
## Attributes Levels PW
## Flavor Chocolate 0.000
## Flavor Mango 1.522
## Flavor Raspberry 0.522
## Flavor Strawberry 0.767
## Flavor Vanilla 1.389
## Packaging Cone 0.000
## Packaging Homemade waffle -0.244
## Packaging Pint -0.100
## Light Low fat 0.000
## Light No low fat 0.478
## Organic Not organic 0.000
## Organic Organic 0.307
## Base utility ~ 4.358
##
## Conjoint importance weights:
## Attributes IW
## Flavor 0.597
## Packaging 0.096
## Light 0.187
## Organic 0.120
##
## Conjoint regression results:
##
## coefficient
## (Intercept) 4.358
## Flavor|Mango 1.522
## Flavor|Raspberry 0.522
## Flavor|Strawberry 0.767
## Flavor|Vanilla 1.389
## Packaging|Homemade waffle -0.244
## Packaging|Pint -0.100
## Light|No low fat 0.478
## Organic|Organic 0.307plot(conjoint)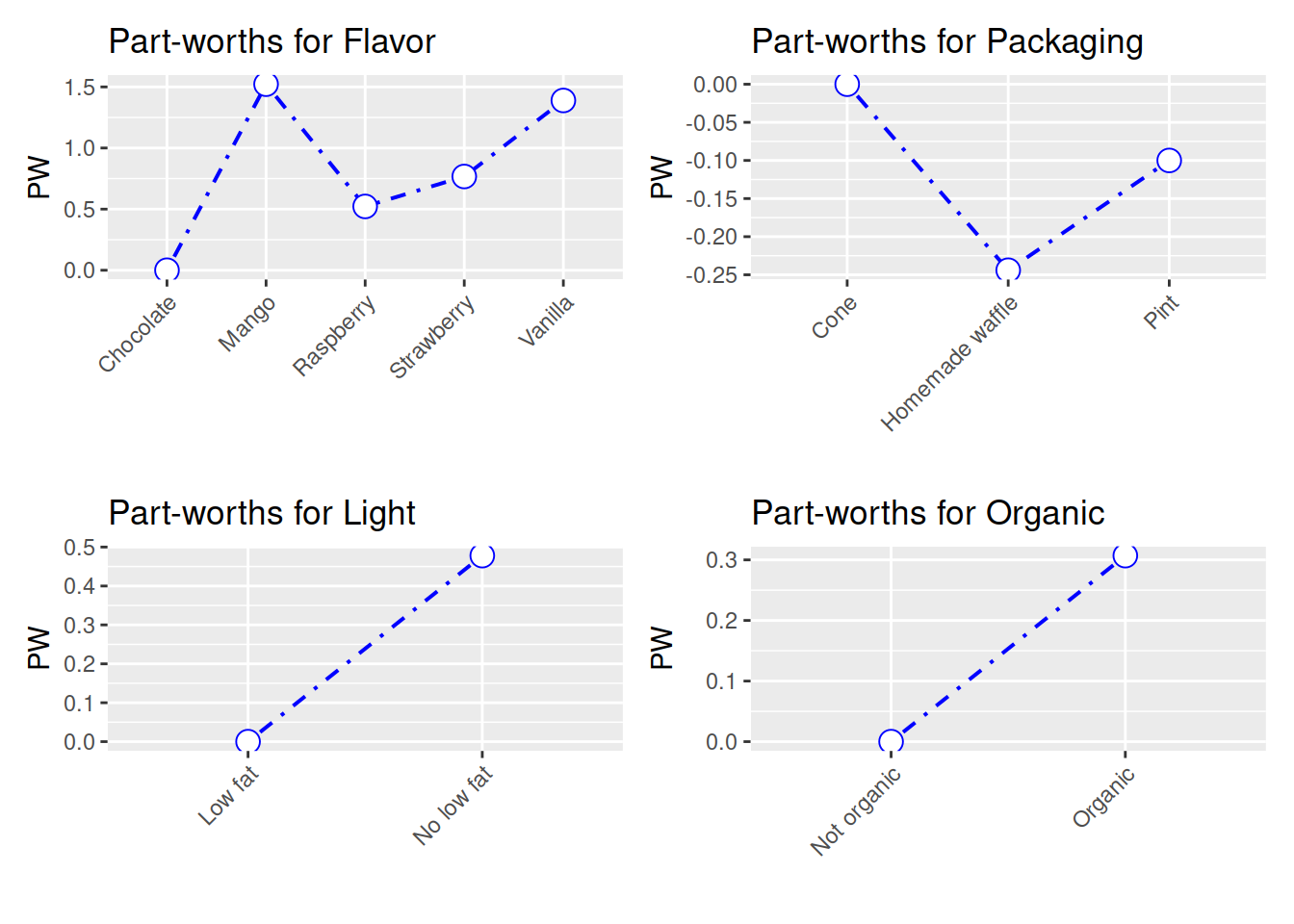 The partworths are just coefficient from a linear regression:
The partworths are just coefficient from a linear regression:
summary(lm(rating ~ Flavor + Packaging + Light + Organic, data = icecream))##
## Call:
## lm(formula = rating ~ Flavor + Packaging + Light + Organic, data = icecream)
##
## Residuals:
## Min 1Q Median 3Q Max
## -5.2867 -2.2244 -0.1911 2.3128 5.4089
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 4.3578 0.8172 5.332 3.76e-07 ***
## FlavorMango 1.5222 0.9453 1.610 0.110
## FlavorRaspberry 0.5222 0.9453 0.552 0.582
## FlavorStrawberry 0.7667 0.8399 0.913 0.363
## FlavorVanilla 1.3889 0.9453 1.469 0.144
## PackagingHomemade waffle -0.2444 0.8675 -0.282 0.779
## PackagingPint -0.1000 0.8399 -0.119 0.905
## LightNo low fat 0.4778 0.5738 0.833 0.406
## OrganicOrganic 0.3067 0.4751 0.645 0.520
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 2.91 on 141 degrees of freedom
## Multiple R-squared: 0.03541, Adjusted R-squared: -0.01932
## F-statistic: 0.6469 on 8 and 141 DF, p-value: 0.7371The conjoint importance weights are the relative importance of each attribute. First the range of partworths for each attribute is calculated. Then the range of partworths for each attribute is divided by the sum of the ranges of all attributes. In this example, Flavor is the most important attribute, with an importance weight of .597.
There is another package called “conjoint” which I have not played with.
In summary, we can use R packages to set up a conjoint experiment, in the sense that we can have partial (fractional) factorial design. After collecting the data, we can use either the conjoint packages, or just run linear or logit models to analyze it. The frequentist approach used in conjoint analysis seems nothing special. Bayesian approach can also be used, but that’s another topic.
Causal effect in conjoint analysis
Hainmueller, Hopkins, and Yamamoto (2014) defined ACME (average marginal component effect). The basic idea is that once all the assumptions are met (usual assumptions in causal inference), we can have the effect of changing one attribute level to another with a causal interpretation. Fortunately it is usually just coefficients from a linear model, or linear combinations of the coefficients. There is a package “cregg” for each calculation and plotting of ACME, but we can just do it manually.